As most of you should know, we’ve been having increased correlated noise in a few sensors around MRP52 (which is disabled) due to it malfunctioning since March 21 2019. This will end with the upcoming service once operations resume, but this post is to clarify how this noise can be cleaned in contaminated data.
A small overview of the issue:
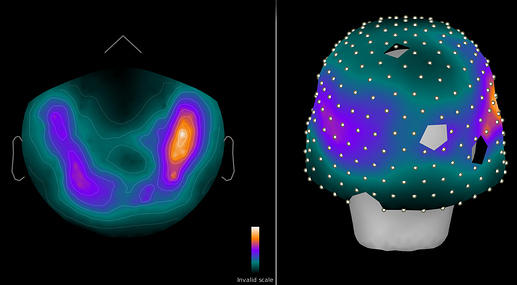
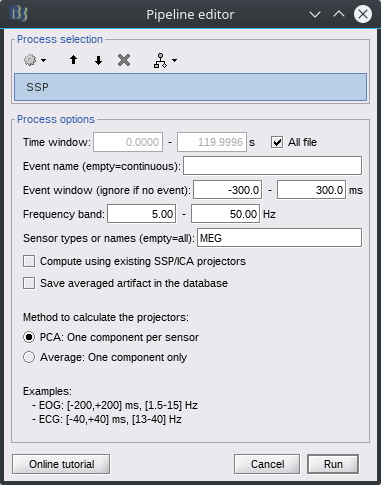
Figure 1: 1-100 Hz power in sensors for resting brain data (pre cleaning)
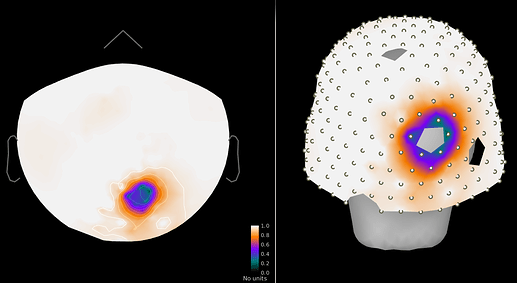
Figure 2: Power ratio post/pre cleaning, ~5-fold drop in closest sensors.
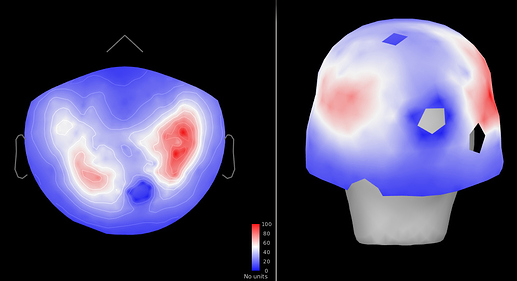
Figure 3: SNR (brain/empty room) (pre cleaning)
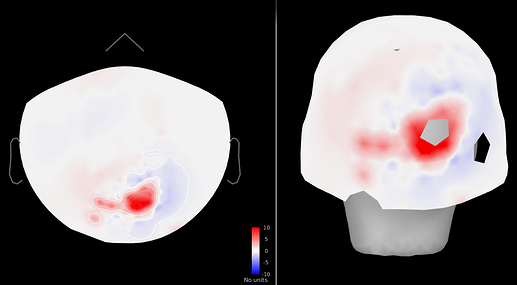
Figure 4: SNR change (post - pre cleaning) ~5-10-fold increase
- Cleaning causes both a large loss in power and a considerable SNR increase in affected sensors.
- SSP cleaning retains some brain signal and is definitely a better alternative to rejecting 5 more neighboring channels.
- However, as the additional noise is not extremely large and would be captured in the noise covariance matrix computed from empty room recordings, cleaning may not be necessary.
Cleaning
Option 1: Do nothing, keeping in mind the increased correlated noise in that area.
Option 2: SSP
- Ensure Brainstorm version is more recent than Apr 2019 (projector import function modified specifically for this)
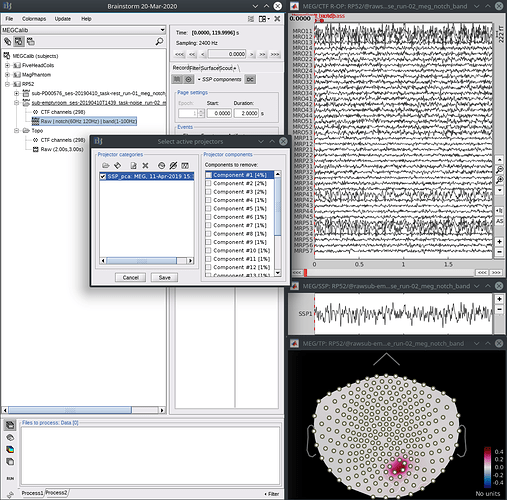
- On the empty room recording, run the generic SSP process. This is a broadband signal, so 5-50 Hz over the entire file works. (Avoid lower frequencies that contain more noise in general.)
Confirm with the topography and/or timeseries that the first component represents the artefact and properly cleans the affected channels.
- From the “select active projectors” panel, save the SSP results to a file using the save icon at the top, right under “Projector categories”.
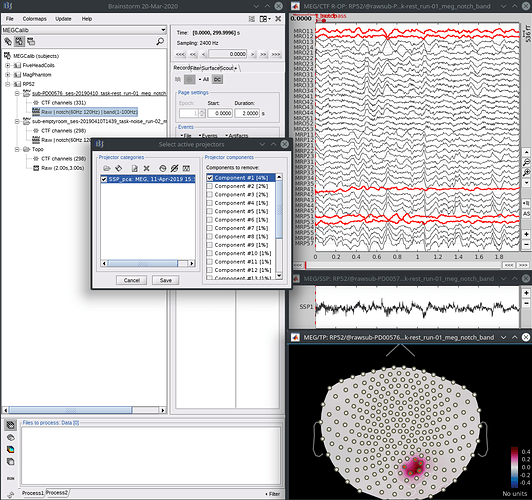
- From a brain recording, open the raw data, then from the “select active projectors” panel, load the projector file you just saved using the load icon (first, top left). Select the first component, see that it reduces both the noise and brain signals in the affected sensors, and save.
I understand both options have down sides. I would recommend doing both and comparing results in a few subjects to decide what’s best for your data.