Table of Contents:
(I) Access to CIVET
(II) Preparing Your Data for Processing
(III) Anonymizing your minc files
(IV) History of Changes in CIVET + Detailed Help
(V) Command-line Interface to CIVET
(VI) CBRAIN Interface:
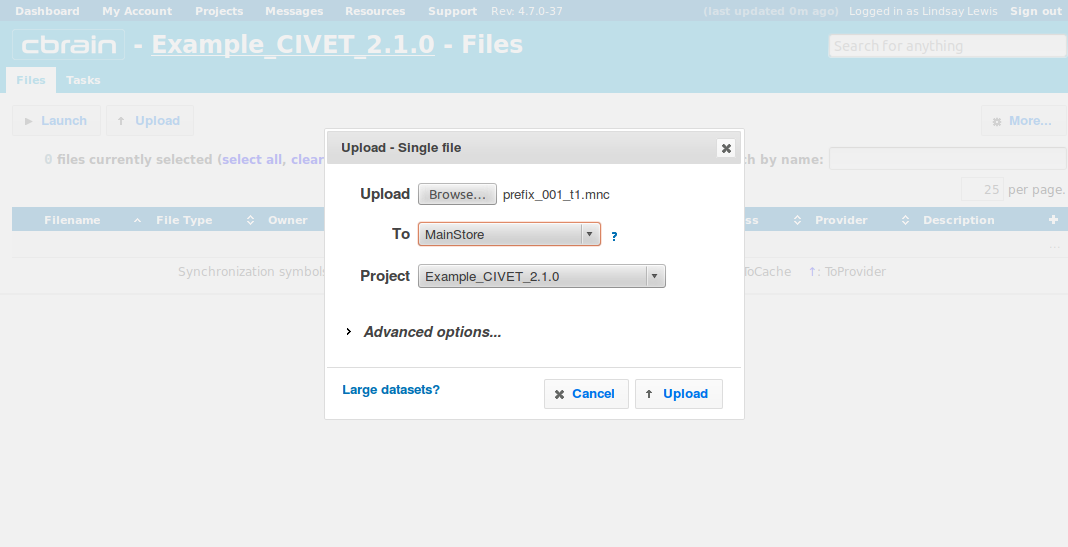
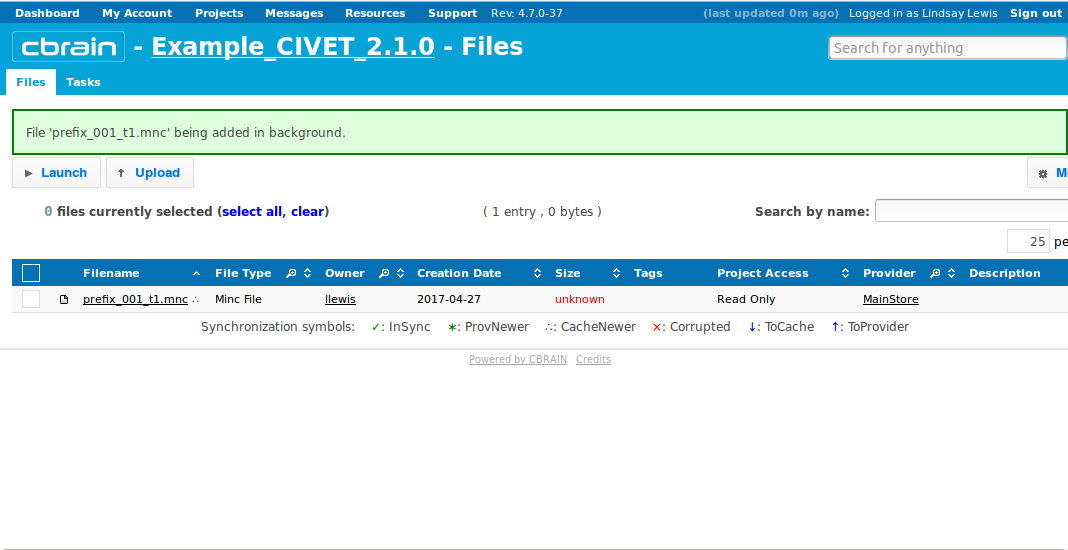
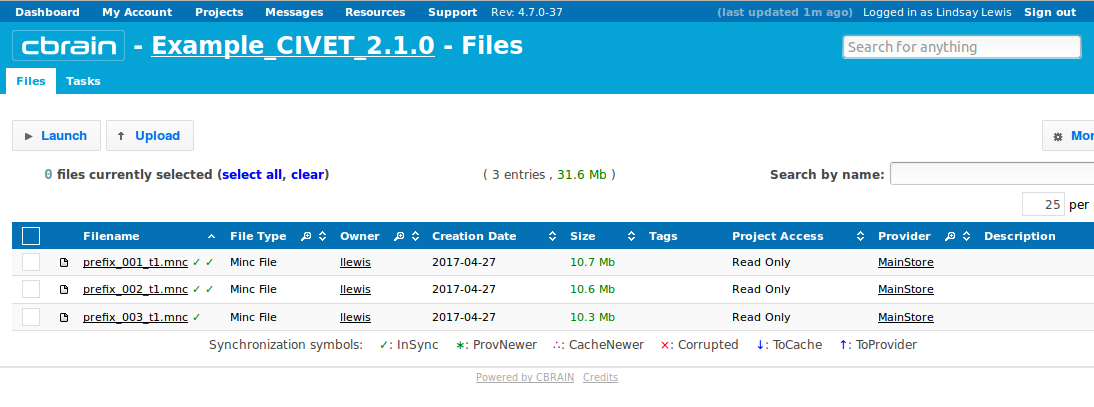
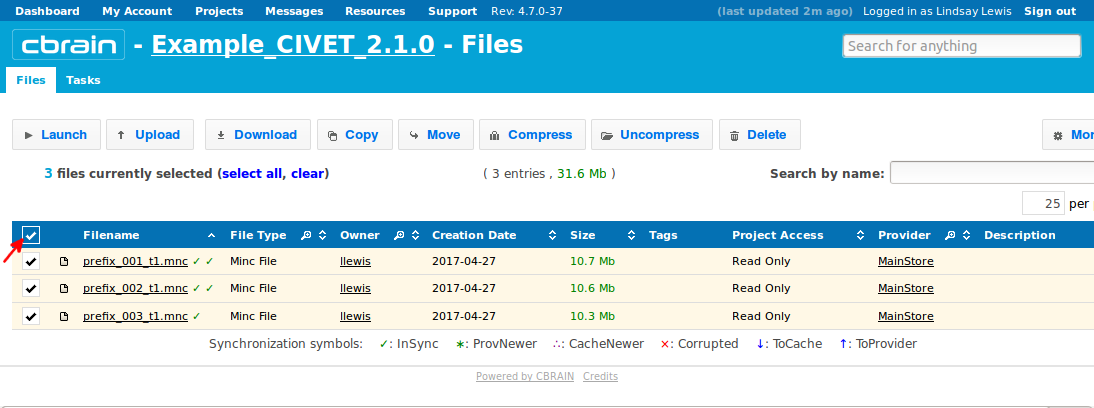
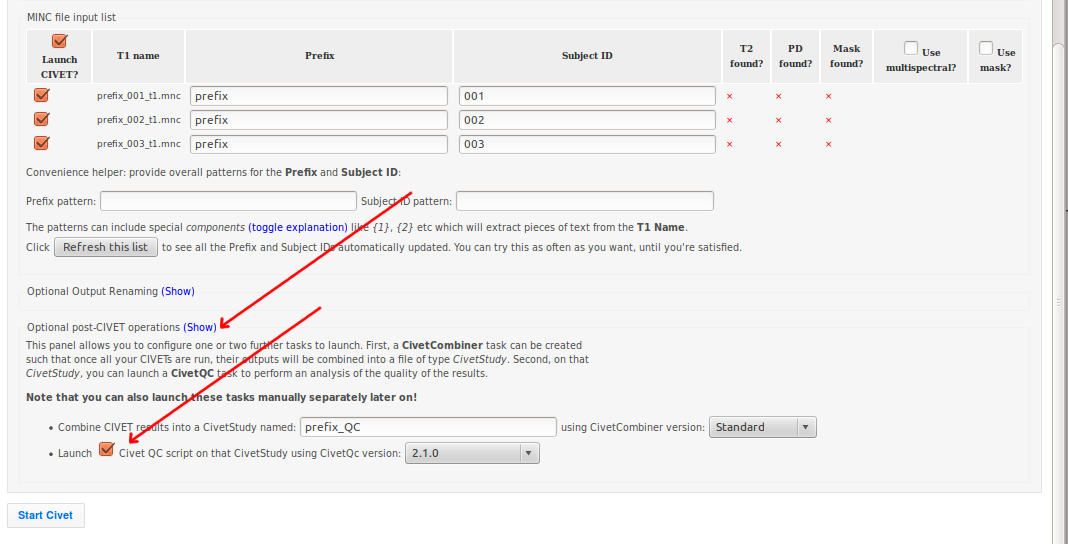
-(1) Inputs
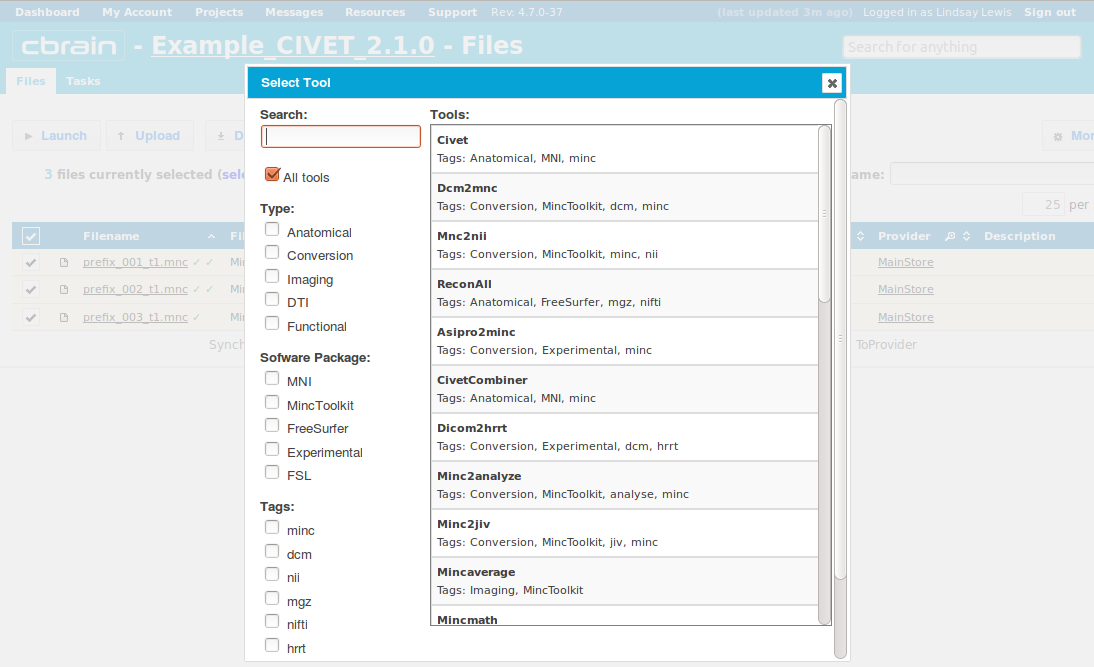
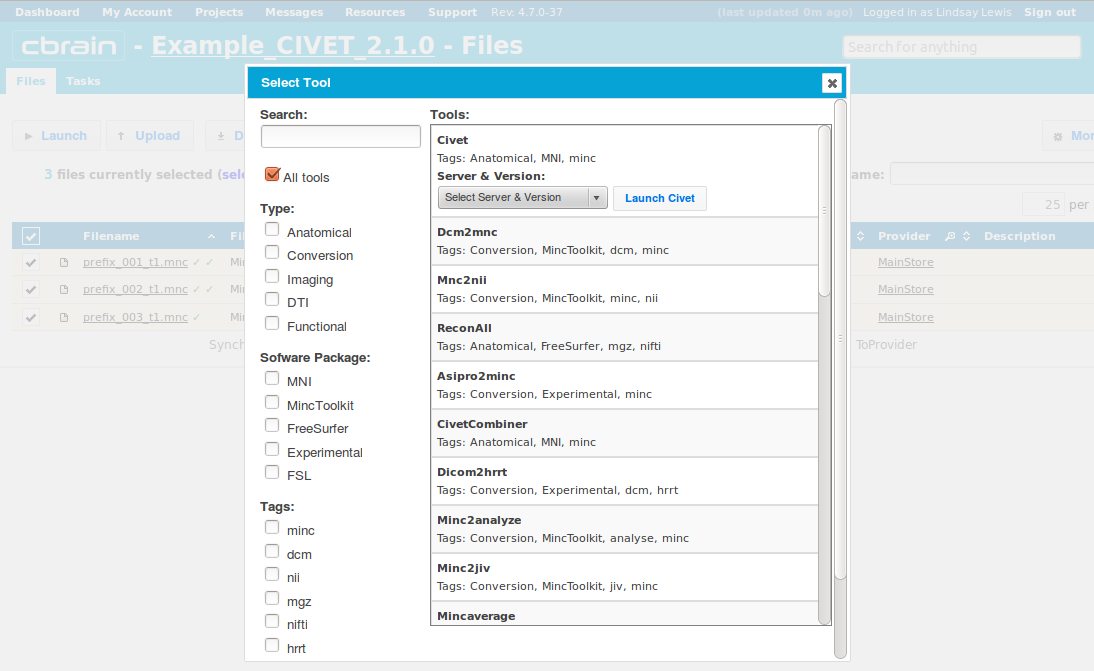
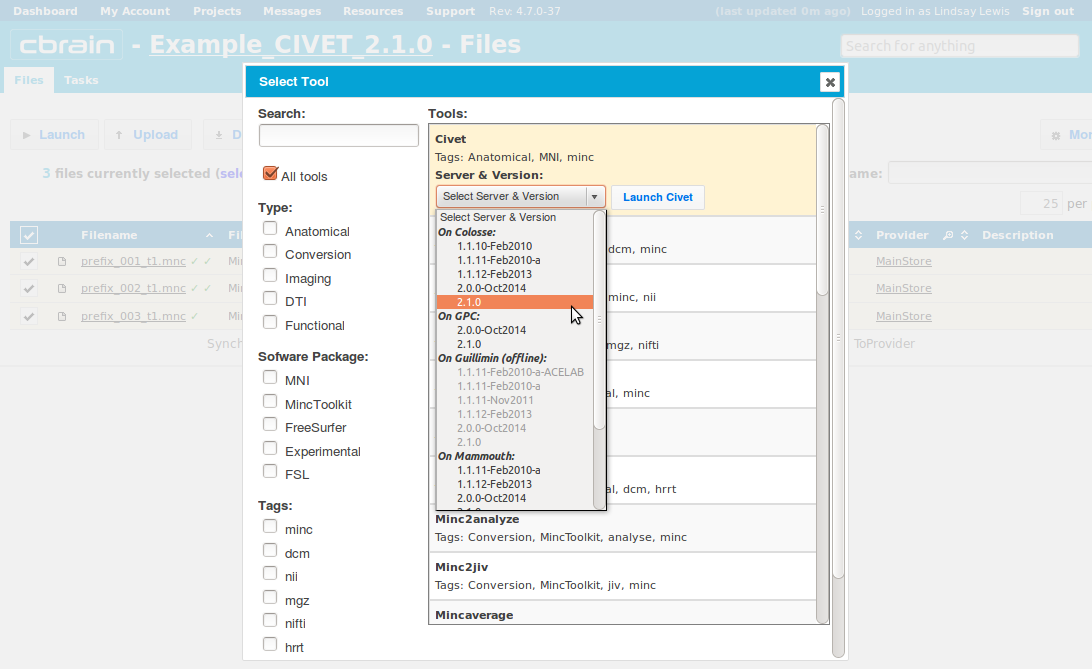
-(2) Choosing a Task
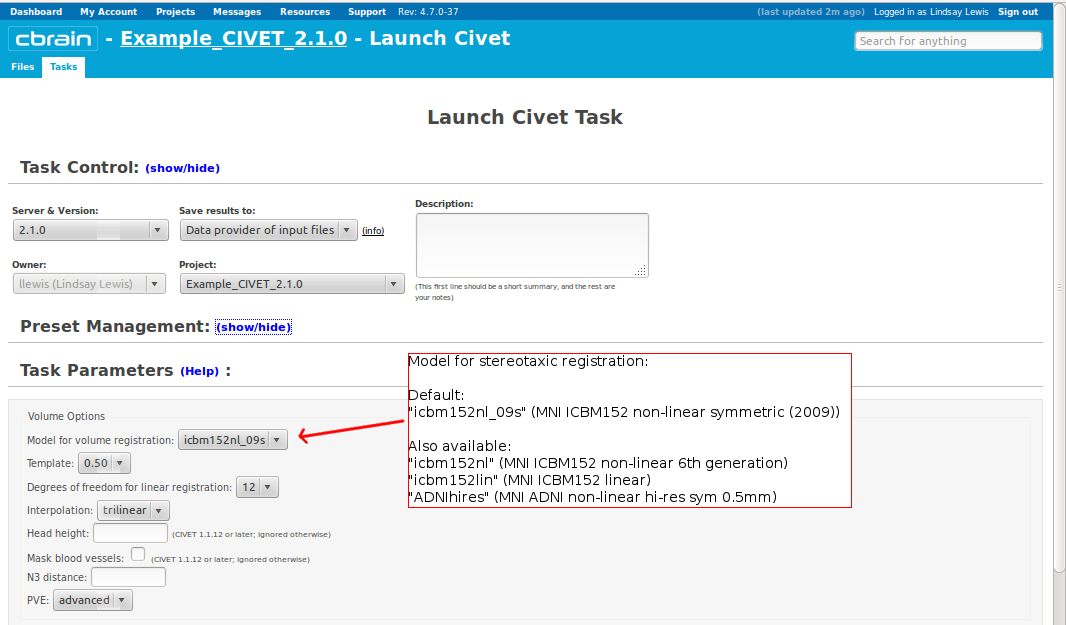
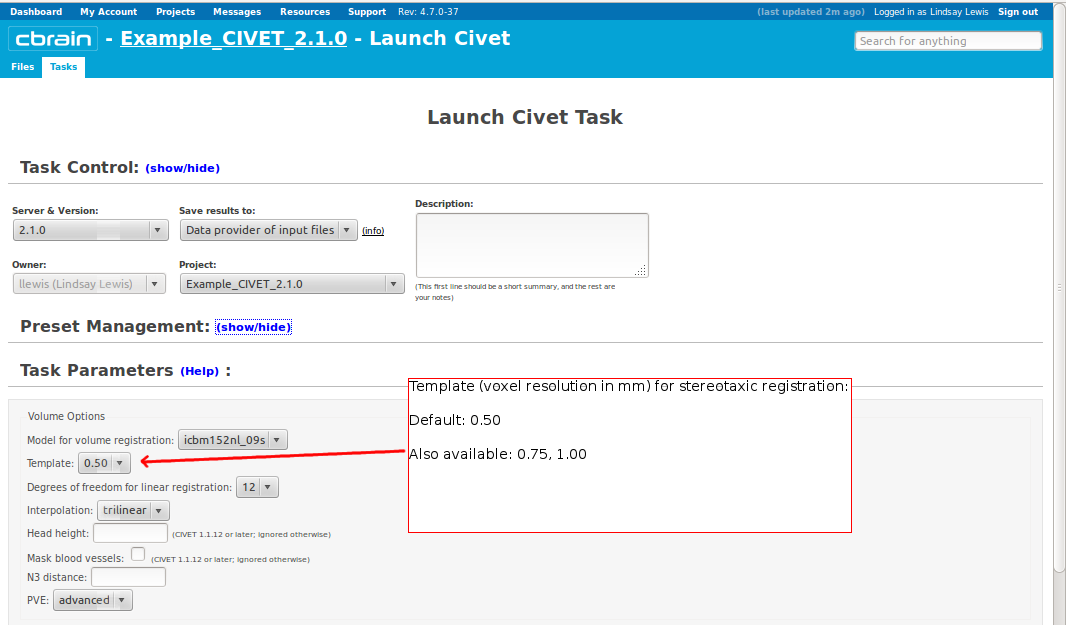
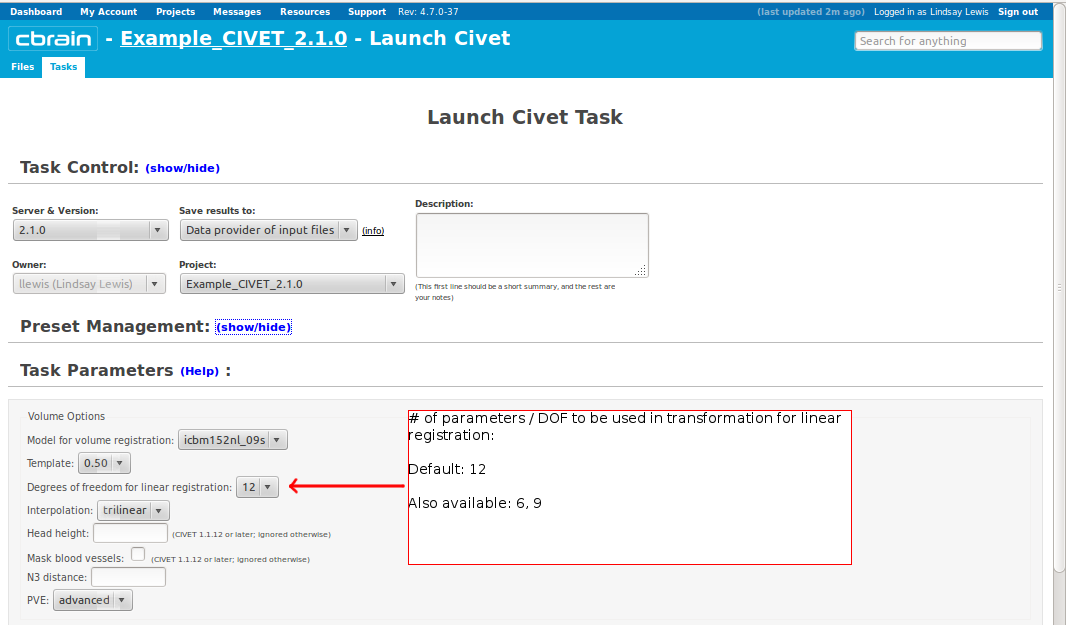
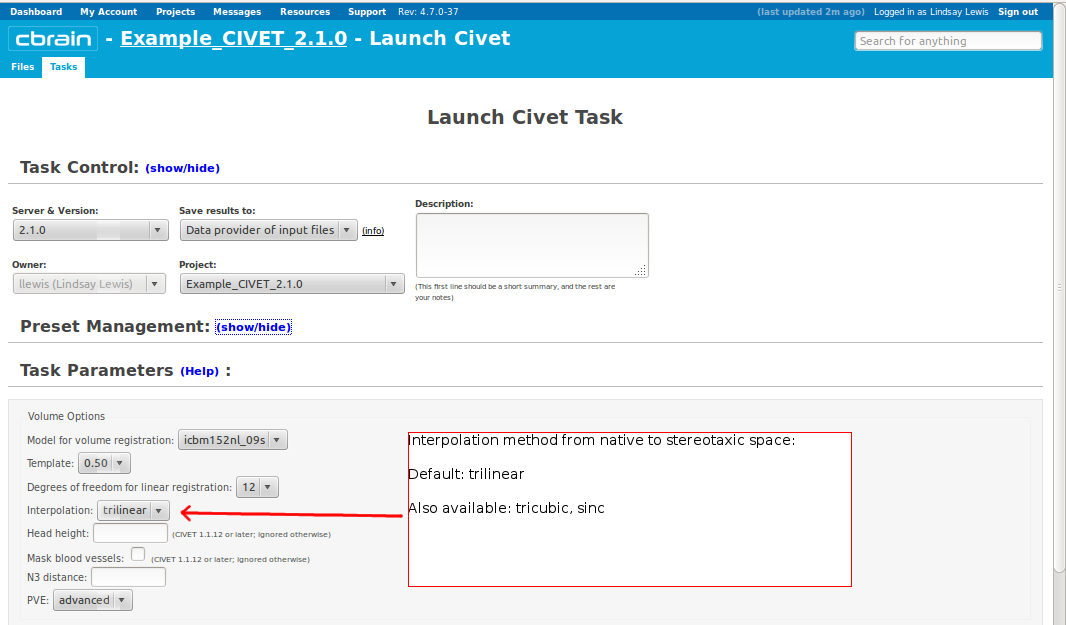
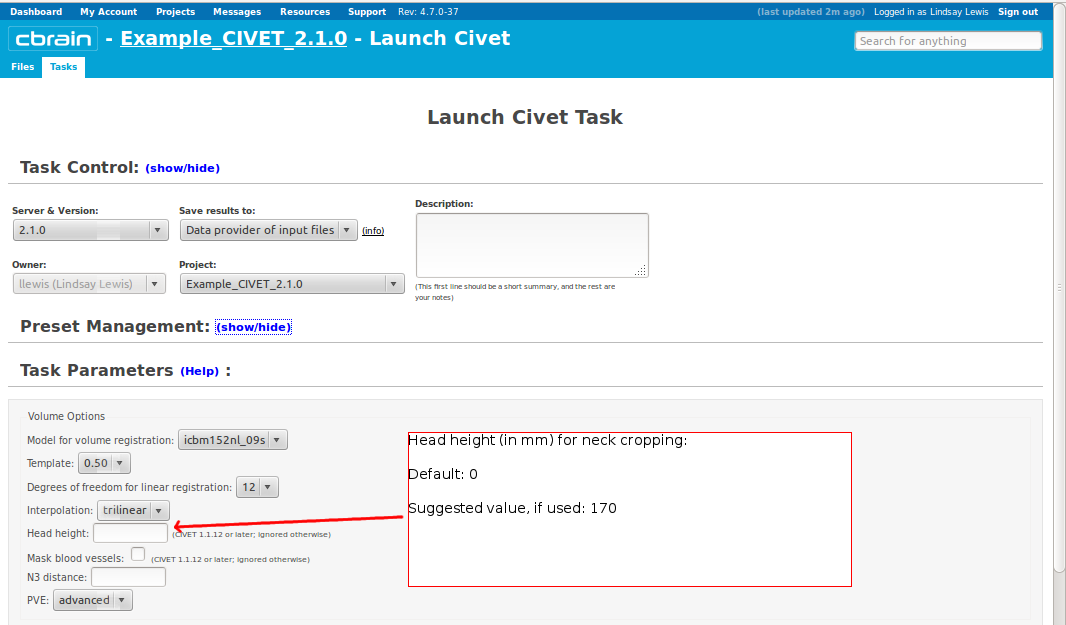
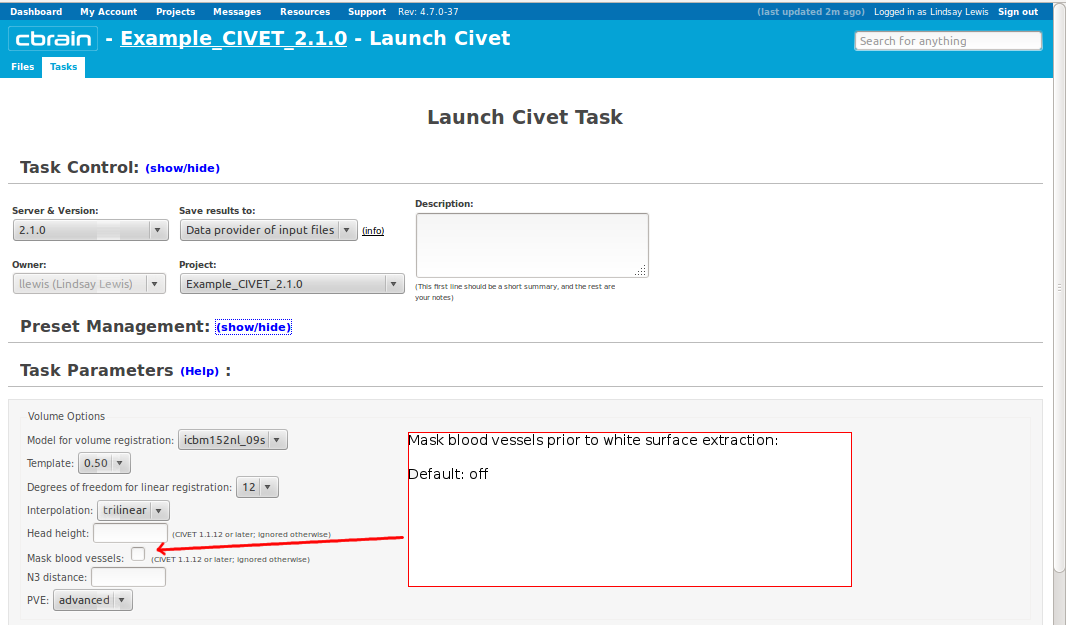
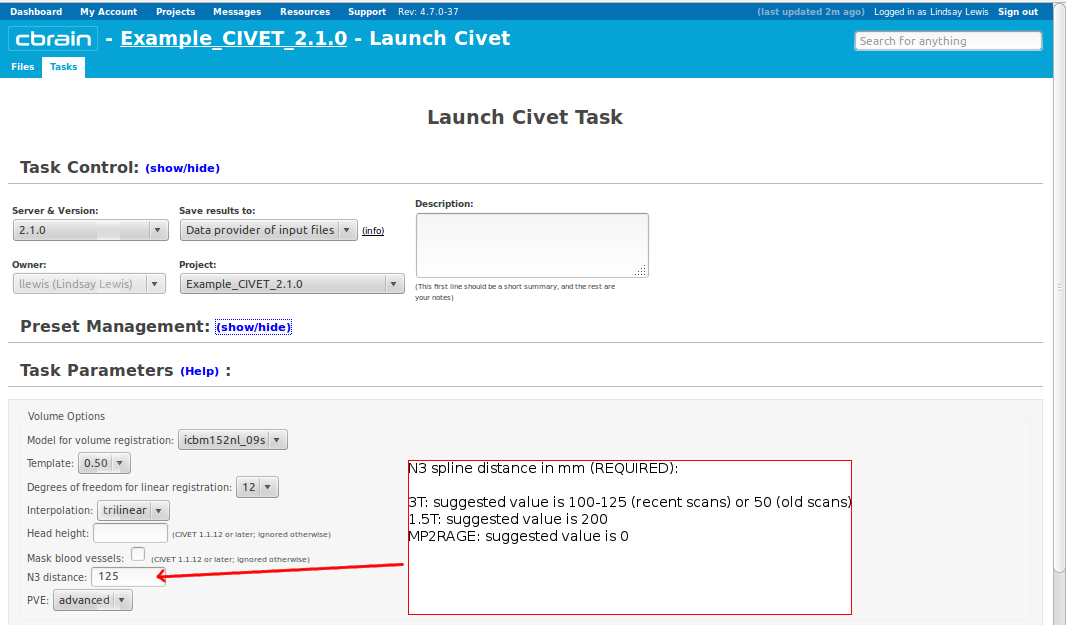
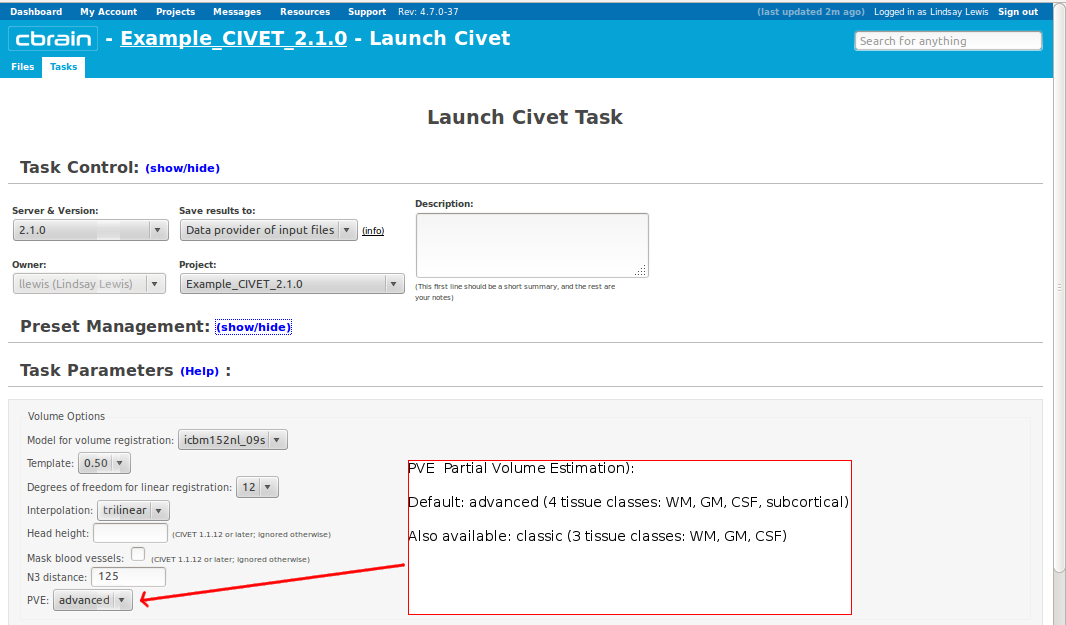
-(3) Volume Options
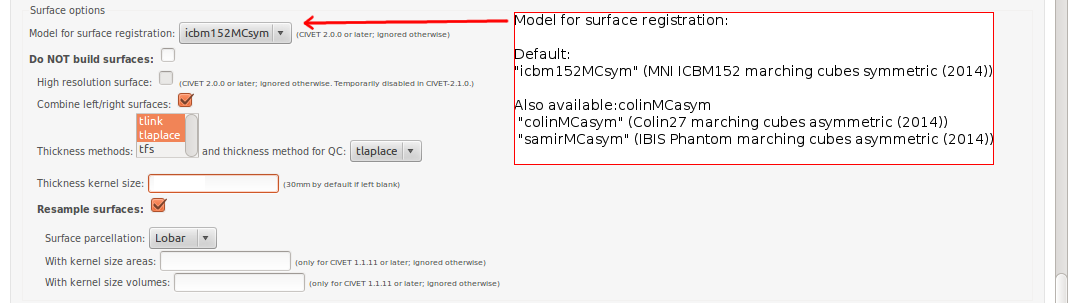
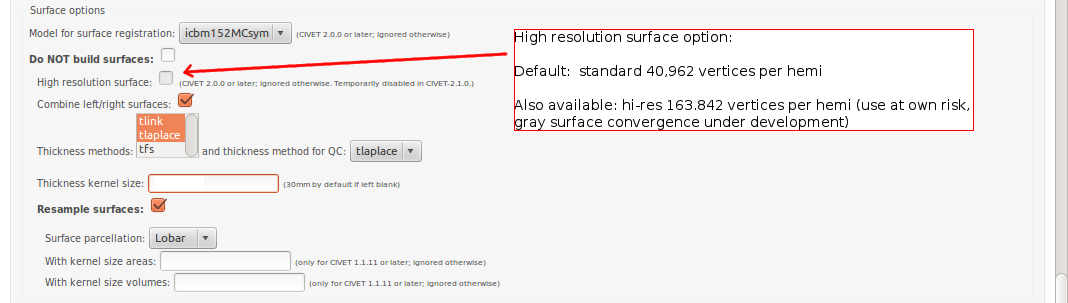
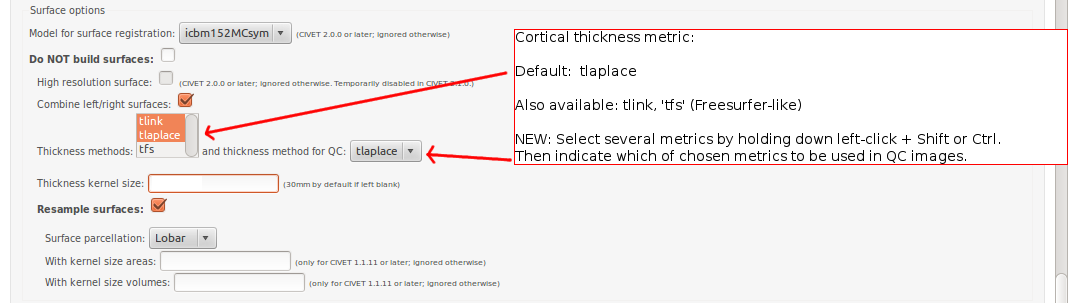
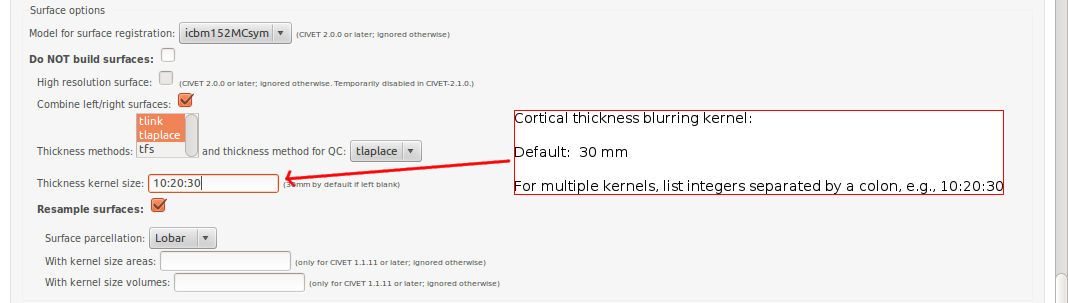
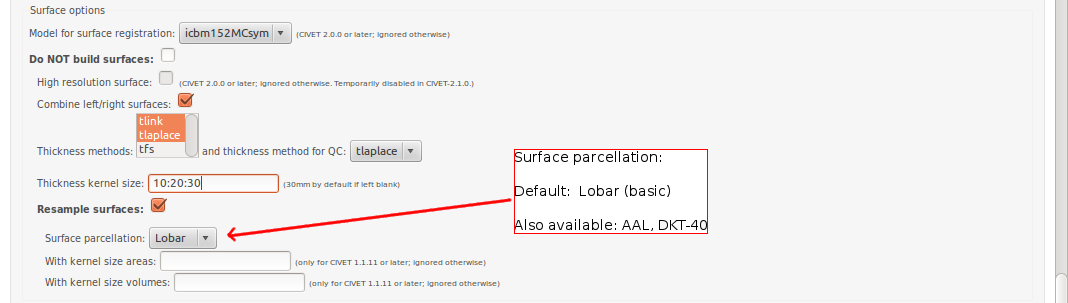
-(4) Surface Options
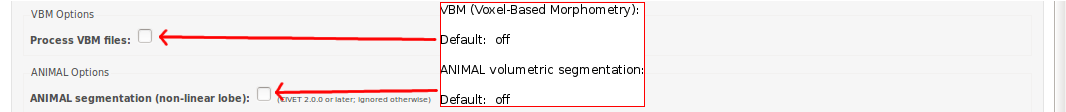
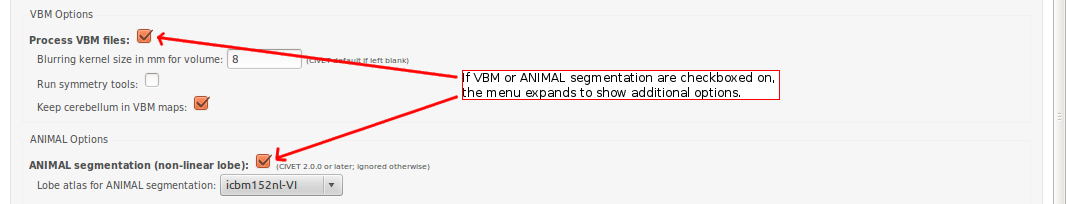
-(5) VBM / ANIMAL Options
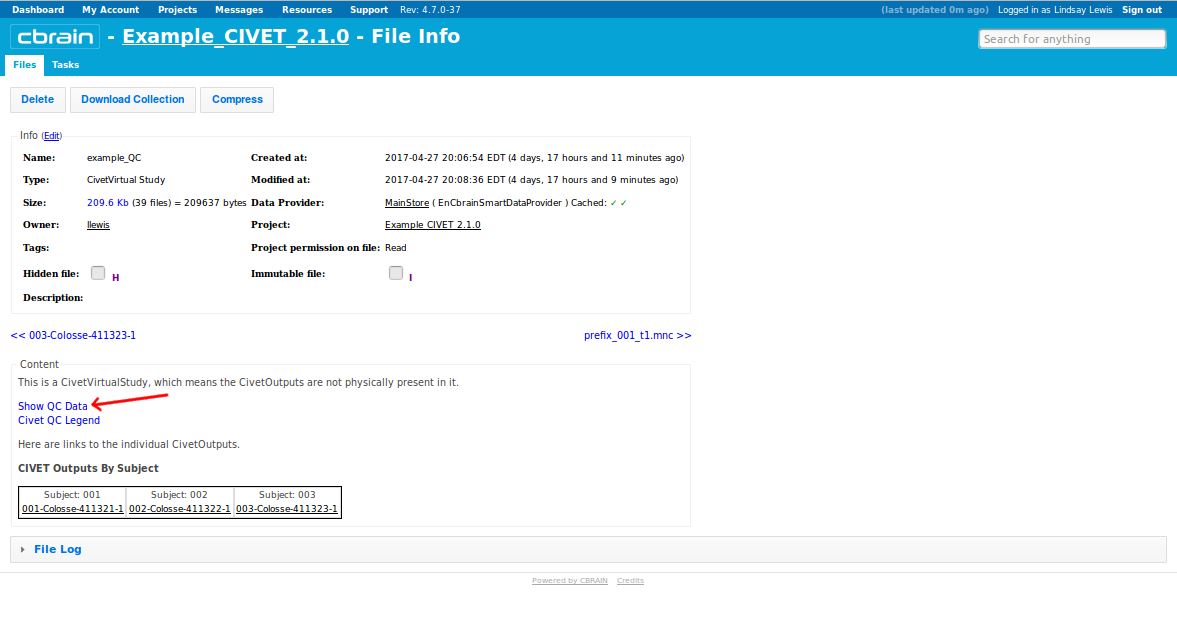
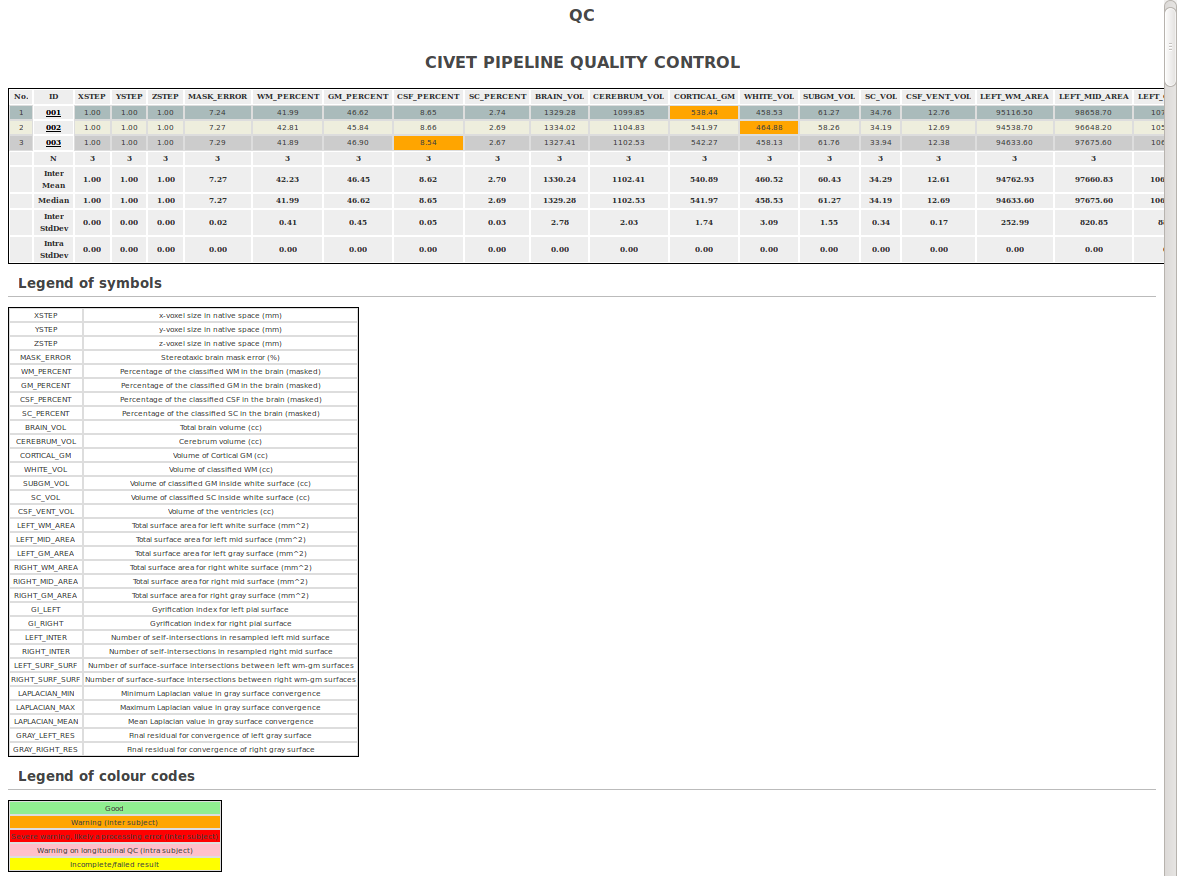
-(6) QC Options
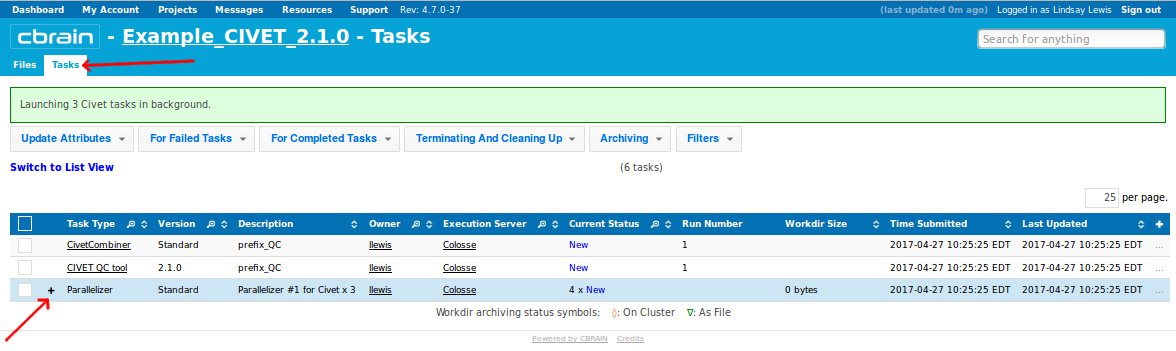
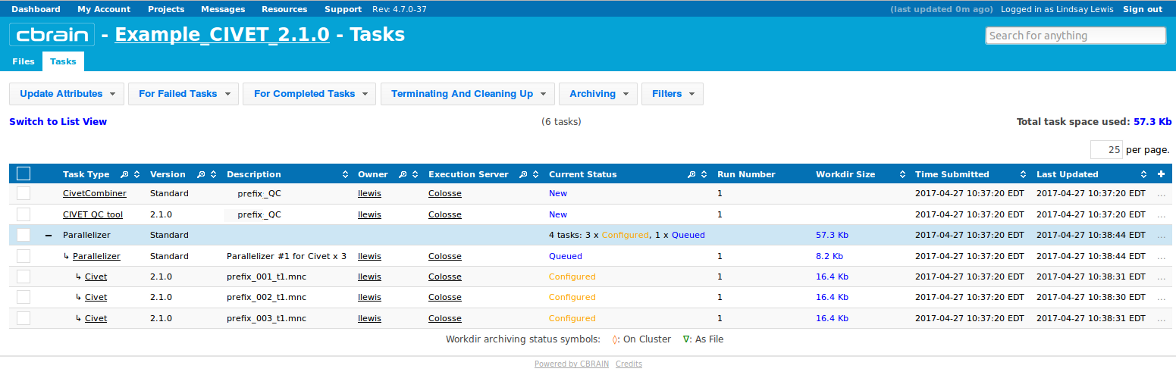
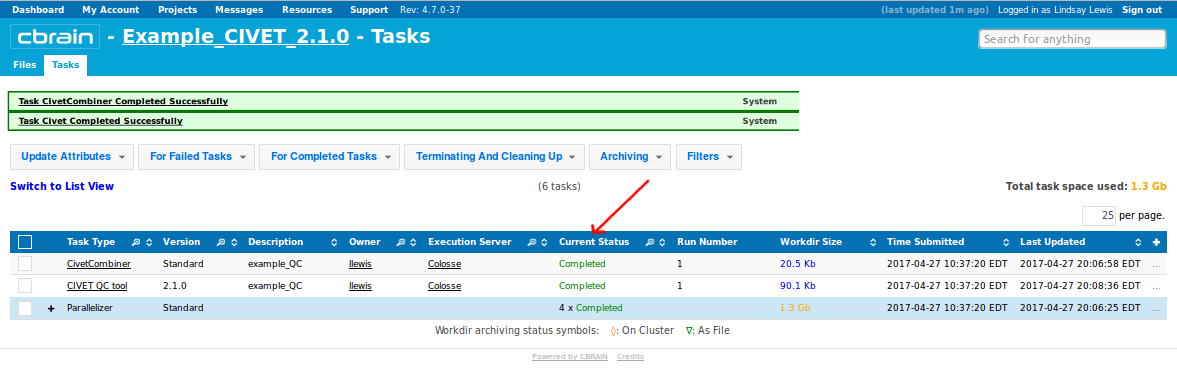
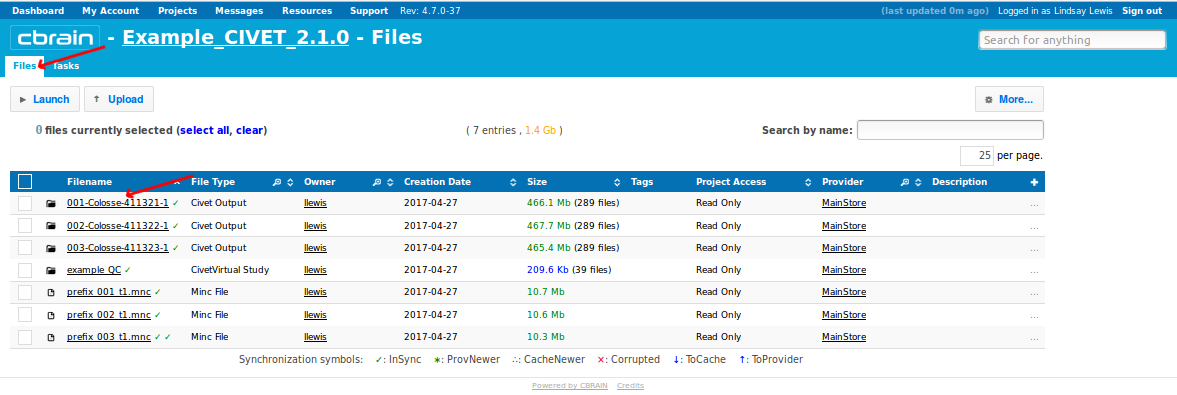
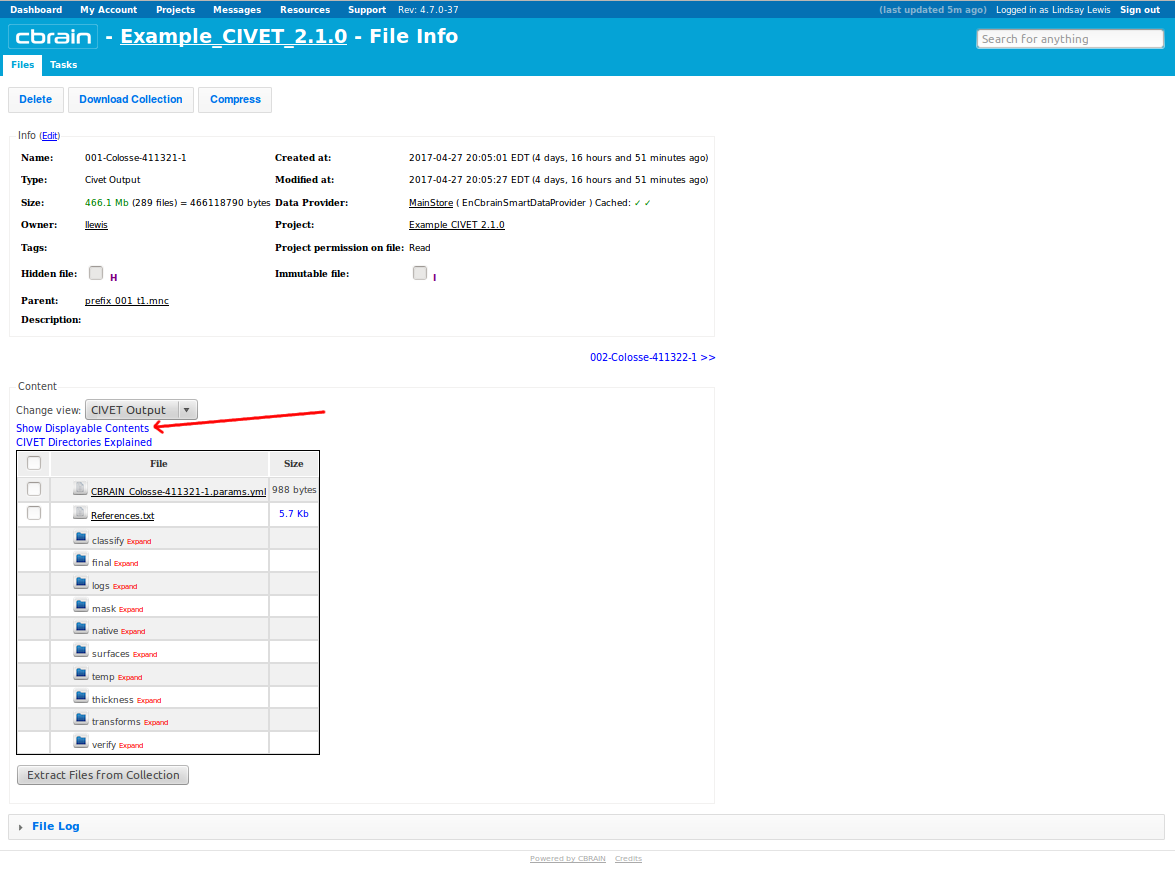
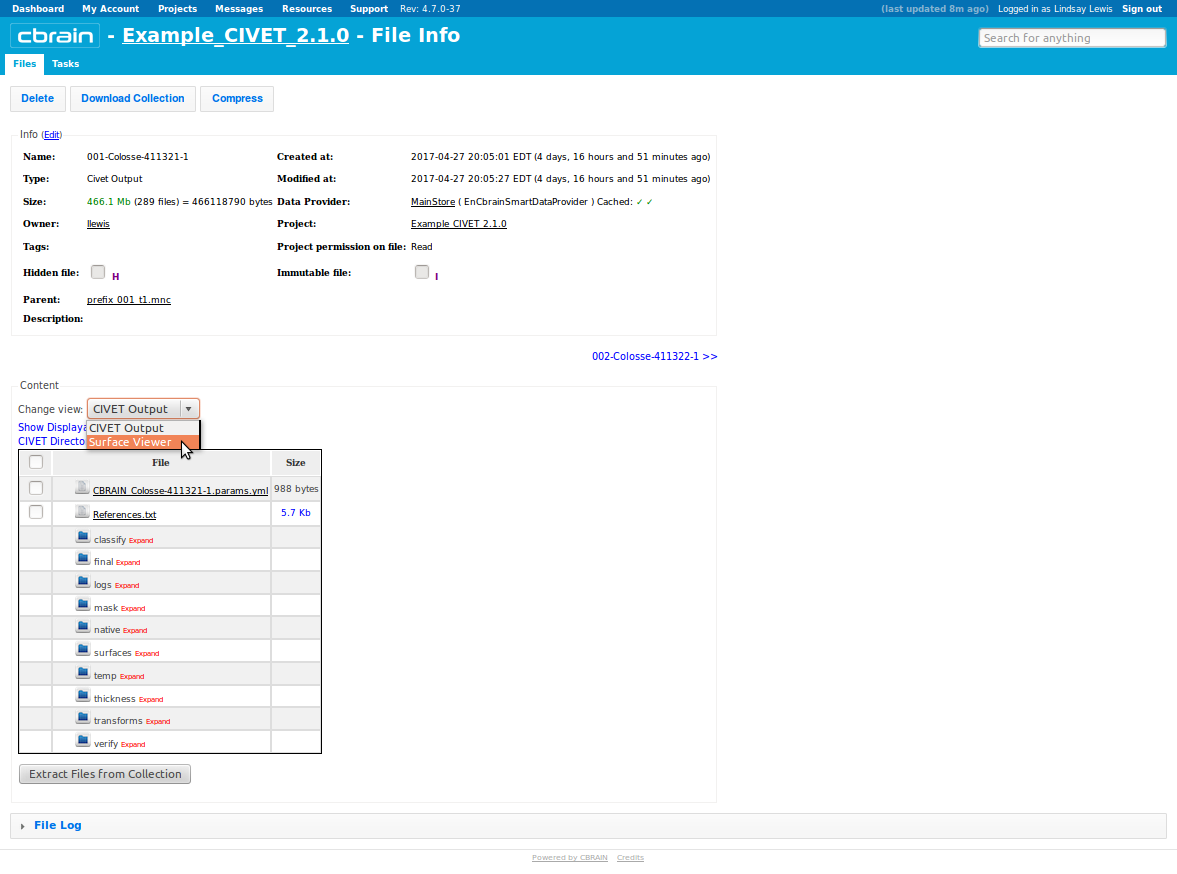
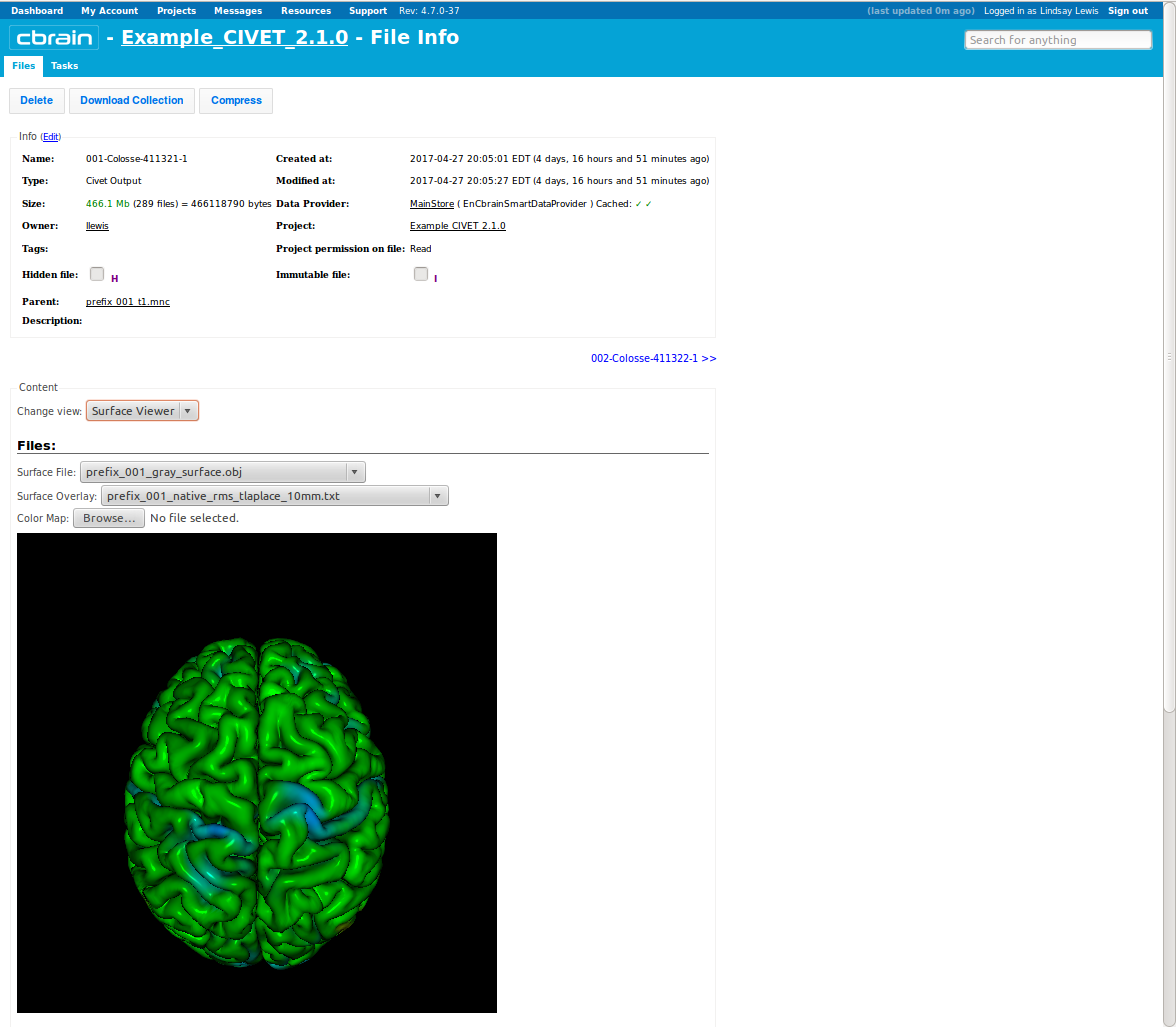
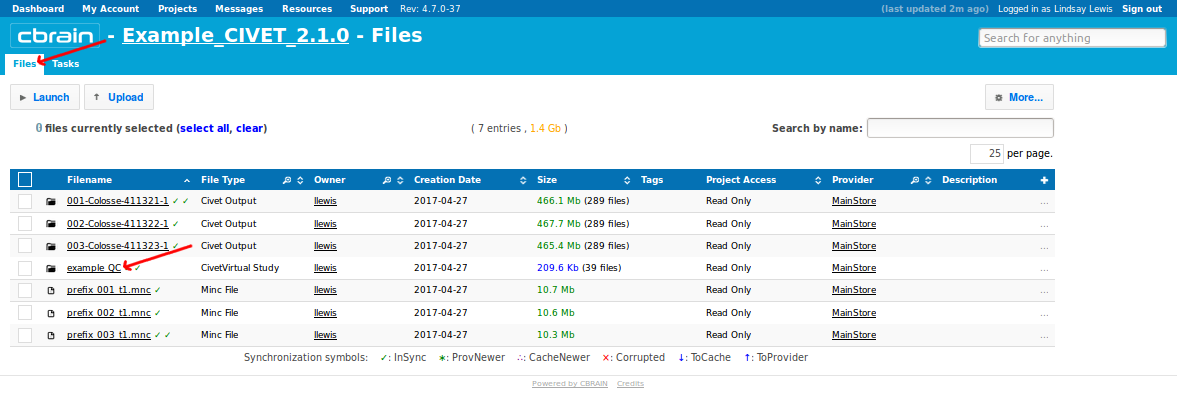
-(7) Viewing your outputs on CBRAIN
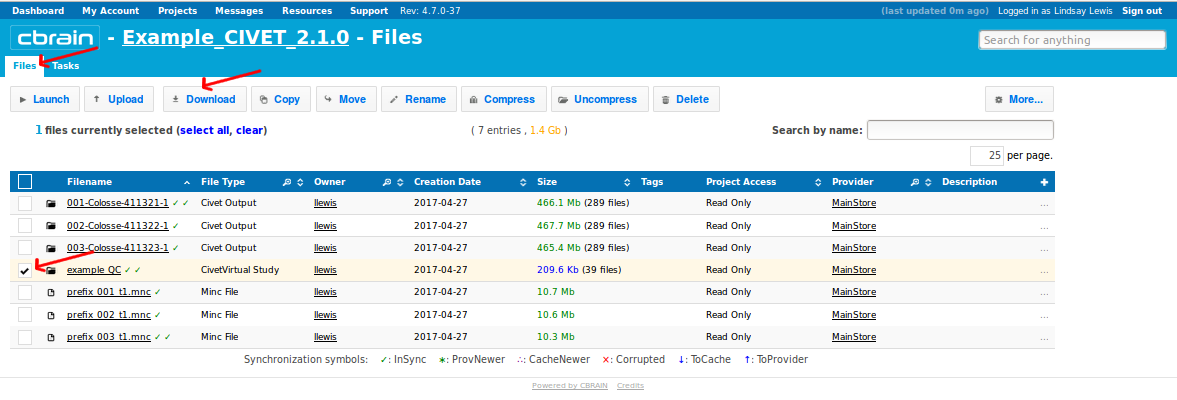
-(8) Downloading your outputs from CBRAIN
Access to CIVET
CIVET is available via the CBRAIN interface, linking high-performance computing (HPC) facilities across Canada and across the world. CBRAIN provides a web-based interface for configuring and running CIVET on large datasets.
Subscription to external users is made possible via collaborative projects. Users are invited to subscribe by providing a description of their project to: cbrain-support.mni at mcgill.ca
Once an account is made, users may access CBRAIN at the CBRAIN portal.
CIVET is no longer supported on the BIC server.
Preparing Your Data for Processing
CIVET requires the image files to follow a strict naming convention. The native MRI files must have the following filename structure:
- $source_t1 =
<prefix>_<id>_t1.mnc - $source_t2 =
<prefix>_<id>_t2.mnc - $source_pd =
<prefix>_<id>_pd.mnc - $source_mask =
<prefix>_<id>_mask.mnc
where $source_t1 is the T1-weighted image file, $source_t2 is the T2-weighted image file (optional), $source_pd is the proton-density-weighted image file (optional), $source_mask is the native brain mask (optional), prefix is the prefix you chose for the project you are working with, and id is the subject identification number. .mnc is the extension that indicates the MINC file format. Images are internally compressed in the MINC2 format to save space.
Images in DICOM, NIfTI, or mgz format can be converted to the MINC format: Converting other file types to MINC. Native MINC files should be pre-processed to make sure they comply to the CIVET pipeline standards: Pre-process MINC files.
It is convenient to keep the source files in a distinct directory. The T1-weighted image must always exist; the T2- and PD-weighted images and mask are optional. Given that most image files come with naming conventions that are very different than the CIVET requirement, we suggest creating a directory that contains symbolic links to the original image files, and use the CIVET naming convention to name the symbolic links.
The T2- and PD-weighted images can be used, if present, to obtain a multi-spectral brain mask and/or for a multi-spectral tissue classification. The input mask, if present, will be transformed to stereotaxic space and be used as a white matter brain mask. The user-supplied brain mask can be useful for eliminating meninges and dura when the automatic masking procedure fails to do so.
Anonymizing your minc files
You may use the following process to make sure that your .mnc files do not contain any sensitive patient information, such as name. You may prefer to script the process if you have a large number of files. Of course, replace firstname and lastname below with your particular subject’s first and last names.
First, if it is not already in the proper (anonymous) naming convention of <prefix>_<id>_t1.mnc , do so now:
mv lastname_firstname_20100610_134507_2_mri.mnc StudyName_01_t1.mnc
Next, search the header for instances of the subject’s name. In this example, we search by first name, but you could equally search by last name.
mincheader StudyName_01_t1.mnc |grep firstname
The output will reveal the fields, if any, where the name is listed.
patient:full_name = “firstname lastname” ; dicom_0×0010:el_0×0010 = “lastname firstname” ; :history = “Fri May 10 11:27:08 2013>>> dcm2mnc -anon -usecoordinates dicom-0005-lastname_firstname_20100610_134507_2_mri <prefix>_<id>_t1
Selectively replace those affected fields with a string (or empty string):
minc_modify_header StudyName_01_t1.mnc -sinsert patient:full_name=‘anonymous’
minc_modify_header StudyName_01_t1.mnc -sinsert dicom_0×0010:el_0×0010=‘anonymous’
minc_modify_header StudyName_01_t1.mnc -sinsert :history=‘’
Search your file again for your subject’s name. This time, it should come up blank.
mincheader StudyName_01_t1.mnc |grep firstname
If desired, you may do the same for age or gender:
minc_modify_header StudyName_01_t1.mnc -sinsert patient:age=‘’
minc_modify_header StudyName_01_t1.mnc -sinsert patient:birthdate=‘’
minc_modify_header StudyName_01_t1.mnc -sinsert patient:sex=‘’
To see a list of all patient fields (with potentially sensitive information), type:
mincheader StudyName_01_t1.mnc |grep patient:
You may also use Andrew Janke’s perl script to remove DICOM headers: minc_de-dicomerise
History of Changes in CIVET + Detailed Help
The -help text in CIVET gives a history of the changes in CIVET over the years and shows all of the available command-line options.
CIVET_Processing_Pipeline, version 2.1.0, released September, 2016. Takes any number of multi or single spectral input MINC volumes and extracts the cortical surfaces from them utilizing the PMP pipeline system. It then calculates cortical thickness at each vertex of the produced cortical surfaces (non-linearly registered) using the t-link metric (in both Talairach and native spaces). It can also produce ANIMAL segmentations, symmetry analyses, regional thickness, surface areas and volumes for brain lobes. On-line documentation is available at: http://www.bic.mni.mcgill.ca/ServicesSoftware/CIVET. ************************************************************************* New in September, 2016 quarantine, version 2.1.0 - allow native scans in stereotaxic space without linear transformation - pve with iterative correction of thresholds, masking of cerebellum and brainstem, sub-cortical tissue class - sub-cortical tissue class to VBM - study prefix is optional - preserve true native space with direction cosines and field of view - compute several thickess methods at once - increase search radius for white surface gradient correction and use Taubin smoothing on white surface - add partial volume estimates to Laplacian field for gray surface expansion - start IBIS v24 model - implement parallelization of subjects via mpirun - QC: add gradient, distortion and convergence graphs - average_surface_builder: allow non-linear mode, add Taubin smoothing ************************************************************************* New in Oct, 2014 quarantine, version 2.0.0 TO DO - Add asym maps for area and volume. 1 - marching-cubes algorithm for extraction of initial white surface 2 - new calibration of white surface at GM-WM t1-gradient 3 - improved node movement in surface fit program 4 - corrected functional for Laplacian constraint in surface fit program (for gray surface) 5 - improved surface registration 6 - new surface registration models based on marching-cubes 7 - simplified CIVET QC pipeline 8 - corrected pve classification at 0.50mm volume template 9 - all changes listed below for 1.1.13 10 - ANIMAL volumetric lobar segmentation 11 - corrected fwhm for surface smoothing (was off by sqrt(2) in 1.1.12) 12 - use grey surface as mask for compute_icbm_vols for cls_volumes.dat (cls_volumes.dat was unmasked in 1.1.12, thus totally wrong). Note: This version requires the package surface-extraction-3.0.2 or higher. More to come soon: 1 - classification and surface extraction in native space. 2 - intermediate model for linear and non-linear volume registration. 3 - Plug-in for alternate population templates 4 - removal of self-intersections in .sm file for surface registration (Maxime Boucher) 5 - Longitudinal surface registration and QC ************************************************************************* New in July, 2013 quarantine, version 1.1.13 (not released) 1 - bug fixed in N3 to be invariant to voxel size (requires N3–1.12.0 and EBTKS-1.6.4) 2 - high-resolution surfaces at 320k polygons 3 - surface registration on high-resolution surfaces, true multi-resolution algorithm (faster) 4 - improved medial cut through the corpus callosum 5 - processing at voxel sizes of 0.5mm and 1.0mm 6 - speed optimizations for pve 7 - speed optimizations for Laplacian field at 0.5mm 8 - cropping of non-linear xfm grid file to save disk space and to reduce peak memory 9 - masking of hippocampus and amygdala for icbm152nl_09s model (0.5mm) 10 - masking of hippocampus and amygdala for ADNIhires model (0.5mm) 11 - improved schedule for white matter surface extraction (smoother convergence, scaled starting ellipsoid) 12 - new ICBM surface average for scaled ellipsoid 13 - clean-up of connected pieces of white matter (partial volumes) 14 - new pipeline to build average surfaces from CIVET output ********** From Feb-2013 quarantine, version 1.1.12 ********** 1 - Neck-cropping option on native image (-headheight) 2 - Improved linear registration (now using normalized mutual information and with model_mask on last fitting stage only) 3 - Improved non-linear registration (now with a headmask and with reduced weight parameter, 2mm steps and higher sampling) 4 - Application of population model’s head mask to reduce background noise in image for mincbet 5 - Final pve classification is unmasked so it can recover any small bits of tissue missing from the brain mask 6 - Improved masking of white matter mask using cortical_surface with a new model (on which the non-linear transform is applied) 7 - Use smoothing from depth_potential instead of diffuse for cortical thickness and surface registration (more accurate smoothing and no more need for BLAS and LAPACK). Note: 20mm with old code is like 30mm with new code. 8 - Improved filling of ventricles using an improved atlas (for white surface extraction) 9 - Fixed WM/GM border by not allowing CSF/BG next to WM (under brainstem, mostly) 10 - New ADNI population models 11 - Generation of basic glim file as part of the CIVET QC 12 - New verify figures for Laplacian fit and surface-surface intersections ********** From Nov-2010 quarantine, version 1.1.11 ********** 1 - option for user-defined surface parcellation atlas. 2 - AAL surface parcellation atlas in models. 3 - normalized cortical volumes maps on resampled surfaces (Lu Zhao and Maxime Boucher). ********** From Feb-2010 quarantine, version 1.1.10 ********** 1 - option for 3 Tesla. 2 - options for reset-after and reset-to (PMP). 3 - cortical area expansion/contraction maps on resampled surfaces. 4 - new ICBM 152 non-linear model (40th generation, 2009a), symmetric and asymmetric. 5 - asymmetry maps for cortical thickness (now normalized) 6 - asymmetry maps for position 7 - basic lobar surface parcellation for quality control of surface registration 8 - sum of absolute mean curvature 9 - gyrification index for whole brain (in addition to by hemisphere) 10 - GI and mean curvature on white, gray, mid surfaces ********** From Feb-14–2008 quarantine, version 1.1.9 ********** 1 - minc2-based quarantine. 2 - new ICBM templates compatible with non-linear ICBM152 model. 3 - asymmetry maps for cortical thickness. 4 - cortical surfaces for hemispheres combined into single surface for convenience of visualization. 5 - integrated VBM pipeline inside CIVET. 6 - list of references produced by CIVET in targetdir. 7 - really mask the background t1-image during classification to avoid confusion between noisy background and csf. 8 - fixed tlaplace cortical thickness for extents of volume in native space. 9 - ANIMAL segmentation only available as a plug-in module. 10 - resampled cortical surfaces in stereotaxic space. 11 - total cerebral volume. ********** From Jul-09–2007 quarantine, version 1.1.8 ********** 1 - ANIMAL segmentation has been disabled as the new default. 2 - recompute non-linear transform to linear symmetric icbm152 model when using the old ANIMAL parcellation. 3 - simplification of PMP options and new -no-granular mode to submit all stages for one subject in one qsub job. 4 - susceptibility artefact map now uses brain with cerebellum. 5 - use a t1-native user-defined custom brain mask, if present, of the form SOURCE_DIR/PREFIX_ID_mask.mnc (or .gz) for linear registration and white matter masking. 6 - use t1only for native brain mask (no need for multispectral mask in native). 7 - run nu_correct in stereotaxic space on the interpolated original native image, not on the interpolated nu_corrected native image. 8 - init environment files are now in the quarantine base directory (no longer in CIVET directory). ********** From previous Mar-30–2007 quarantine, version 1.1.7 ********** 1 - added mean curvature on native mid surfaces. 2 - added gyrification index. 3 - cortical thickness now always produced, with tlink method and 20mm blurring kernel as defaults. 4 - added percentages for tissue types in verify image. 5 - much improved brain mask, remove -crop-neck option. ********** From previous Mar-20–2007 quarantine, version 1.1.6 ********** 1 - support for various interpolation methods from native to stereotaxic (linear, cubic, sinc; default is linear). 2 - error metric for brain mask in native space (in verify image). 3 - shuffle the PMP steps so that -resetAll actually works. 4 - file name extension for surface maps is .sm, not .mnc. (use rename ‘s/.mnc$/.sm/’ /transforms/surfreg/.mnc). 5 - can use a 0mm blurring kernel for cortical thickness (no blurring). 6 - improved cut through callosum to split hemispheres. 7 - fixed bug in surface registration for infinite loop (on RedHat). 8 - clear history in minc files to go around likely bug in netCDF 3.6.1. ********** From previous Feb-28–2007 quarantine, version 1.1.5 ********** 1 - surface registration and resampling of cortical thickness. 2 - binary masks in byte format to save disk space. 3 - specification of data-processing template for voxel size (0.50mm, 0.75mm, 1.0mm, 2.0mm, etc; default is 1.0mm). 4 - check for irregular slice spacing in native images. 5 - summary of options printed in logs directory of subject. 6 - validation of input parameters for correct values. 7 - possibility to easily rerun CIVET with different options for cortical thickness method and fwhm. 8 - keep mid-surfaces in stereotaxic space 9 - keep surface lobes parcellation at vertices ********** From previous Jan-24–2007 quarantine, version 1.1.4 ********** 1 - add surface segmentation based on ANIMAL’s volume segmentation and calculation of cortex area for lobes (eventually will use surface registration with surface parcellation). 2 - iterative correction to the mean and variance of the tissue types inside pve as the process converges (option -correct-pve) ********** From previous Jan-15–2007 quarantine, version 1.1.3 ********** 1 - add -nothreshold to mritoself for -spectral_mask. This affects only the multispectral mask, in very rare cases. If the native mask looks wrong and linear registration for no obvious reason, with -spectral_mask, then re-run your subject with this fix. 2 - add -lsq6 to bestlinreg.pl and apply initial transform, if given, to mask, only if the mask is given too (does not apply to CIVET). ********** From previous Dec-20–2006 quarantine, version 1.1.2 ********** 1 - use the icbm152 mask for nu_correct instead of the avg305 mask, since subject is linearly registered to icbm152, not avg305. 2 - transformation of surfaces to native space, with cortical thickness evaluated in native space. 3 - quarantine based on minc-1.5 (upgraded versions of pcre, pcre++, CGAL,BOOST for compilation in 64 bits). 4 - use icbm152 surface mask for verify image (red lines). Was avg305 before. 5 - use classified image from pve instead of classify_clean as input to cortical_mask for white matter masking. 6 - improved mincbet for hyperintense voxels in t1 for masking meninges 7 - verify.png now shows, in the first row, the brain mask that was used for linear registration (linear registration can live with a mask that is slightly off, but not too wrong however) 8 - make all.q the default queue instead of aces.q 9 - new flag -lsq12 to CIVET for 12-param linear registration (default is still 9-param) 10 - multispectral classification now produces the good result ********** From previous Nov-02–2006 quarantine, version 1.1.0 ********** The major flaws in the Sep-12–2006 quarantine have been discovered and fixed and the former quarantine should no longer be used for any reason. The Sep-12–2006 quarantine would give too much white matter, thus bad white surfaces and consequently wrong cortical thickness. All simulations done with the Sep-12–2006 quarantine should be redone using the current one. 1 - nu_correct must be performed in Talairach space after linear registration, where a suitable brain mask is available. 2 - multispectral brain mask now available in native space (was available only in Talairach space before) 3 - fixed minc interface to mincbet to copy direction cosines of axes (was responsible for small rotation of brain in Talairach) 4 - linear registration is more robust and is now really “best” 5 - application of the non-linear transformation to the tag points used by classify_clean (essential for young children with small ventricles or AD subjects with large ventricles) There are several other minor enhancements that will not change the results but that will make your like easier: 1 - verify.png image has an outline for the brain mask in Talairach so that you can easily see if the linear registration has been successful 2 - clean-up of the lock files when CIVET is interrupted 3 - source and target directories may use relative paths 4 - source files may or may not be gzipped. *************************************************************************
Summary of options:
— Execution control -----------------------------------------------------------
-spawn Use the perl system interface to spawn jobs [default: use local host scheduler RQCHP] -queue <queue> Which queue to use [default: qwork@ms] -hosts <hosts> Colon separated list of hosts -qopts <opts> Extra options to queuing system [default: “-l walltime=20:00:00 -l nodes=1:ppn=1″] -no-granular Granularity level for submission of jobs using queueing system. [default] -granular opposite of -no-granular -maxqueued <val> Maximum number of jobs that can be submitted at once. [default: 800] -mpi Submit jobs using mpirun to fill all processors on core. -no-mpi opposite of -mpi [default]
— File options ----------------------------------------------------------------
-sourcedir <dir> Directory containing the source files. -targetdir <dir> Directory where processed data will be placed. -prefix <prefix> File prefix to be used in naming output files. -id-subdir Indicate that the source directory contains sub-directories for each id -id-file <file> A text file that contains all the subject id’s (separated by space, tab, return or comma) that CIVET will run on.
— Pipeline options ------------------------------------------------------------
-template <val> Define the template for image processing in stereotaxic space (0.50, 0.75, 1.00, 1.50, 2.00, 3.00, 4.00, 6.00). [default: 1.00] -model <model> Define the model for image-processing: “colin27″ (MNI Colin27 asymmetric (2009)), “icbm152nl_09s” (MNI ICBM152 non-linear symmetric (2009a)), “icbm152nl” (MNI ICBM152 non-linear 6th generation), “icbm152lin” (MNI ICBM152 linear), “ibis-v24″ (MNI IBIS 24 months, symmetric (2015)), “ADNIhires” (MNI ADNI non-linear hi-res sym 0.5mm) [default: icbm152nl_09s] -surfreg-model <model> Define the model for surface registration: “icbm152MCsym” (ICBM152, marching-cubes, symmetric (2014)), “colinMCasym” (Colin, marching-cubes, asymmetric (2014)), “samirMCasym” (IBIS Phantom (Samir), marching-cubes, asymmetric (2014)) -surface-atlas <model> Define the atlas for surface parcellation: “lobes” (coarse lobar parcellation, symmetric), “AAL” (AAL parcellation, asymmetric, based on Colin brain), “DKT” (DKT-40 parcellation, asymmetric) [default: lobes]
— CIVET options ---------------------------------------------------------------
-input_is_stx Assume that the input volume is already linearly regsieted to stx space; this skips the linear registration steps -noinput_is_stx opposite of -input_is_stx [default] -multispectral Use T1, T2 and PD native files for tissue classification. -correct-pve Apply correction to the mean and variance of tissue types at pve iterations. [default] -no-correct-pve opposite of -correct-pve -mask-cerebellum mask cerebellum and brainstem from pve classification [default] -no-mask-cerebellum opposite of -mask-cerebellum -subcortical create a sub-cortical SC class in pve classification [default] -no-subcortical opposite of -subcortical -calibrate-white Apply gradient intensity correction for calibration of white surface. [default] -no-calibrate-white opposite of -calibrate-white -spectral_mask Use T1, T2 and PD stereotaxic files for brain masking. -interp <method> Interpolation method from native to stereotaxic space (“trilinear”, “tricubic”, “sinc”) [default: trilinear] -headheight <dist> head height in mm for neck cropping (use 0 for none). [default: 175] -N3-distance <dist> N3 spline distance in mm (suggested values: 200 for 1.5T scan; 50 for 3T scan). -N3-damping <lambda> N3 damping coefficient (lambda) (suggested values: 2.0e-06). [default: 2.0e-06] -lsq6 use 6-parameter transformation for linear registration [default -lsq9] -lsq12 use 12-parameter transformation for linear registration [default -lsq9] -no-surfaces don’t build surfaces -hi-res-surfaces build high resolution surfaces -mask-blood-vessels mask blood vessels prior to white surface extraction -no-mask-blood-vessel opposite of -mask-blood-vessels [default] -mask-hippocampus mask hippocampus and amygdala for surface extraction if model supports it [default] -no-mask-hippocampus opposite of -mask-hippocampus -thickness compute cortical thickness and blur [tlink][:tlaplace][:tfs] [fwhm1][:fwhm2]:…[:fwhmN] kernel sizes in mm [default: tlink 30] -resample-surfaces resample cortical surfaces -no-resample-surfaces opposite of -resample-surfaces [default] -mean-curvature produce mean curvature maps on surfaces -no-mean-curvature opposite of -mean-curvature [default] -area-fwhm <fwhm> fwhm1:fwhm2:…:fwhmn blurring kernel sizes in mm for resampled surface areas [default: 40] -volume-fwhm <fwhm> fwhm1:fwhm2:…:fwhmn blurring kernel sizes in mm for resampled surface volumes [default: 40] -combine-surfaces combine left and right cortical surfaces -no-combine-surfaces opposite of -combine-surfaces [default]
— VBM options -----------------------------------------------------------------
-VBM process VBM files for analysis [default -no-VBM] -no-VBM don’t process VBM files for analysis -VBM-fwhm <fwhm> blurring kernel size in mm for volume [default: 8] -VBM-symmetry run symmetry tools [default -no-VBM-symmetry] -no-VBM-symmetry don’t run symmetry tools -VBM-cerebellum keep cerebellum in VBM maps -no-VBM-cerebellum mask out cerebellum in VBM maps [default -VBM-cerebellum]
— ANIMAL options --------------------------------------------------------------
-animal run volumetric ANIMAL segmentation [default -no-animal] -no-animal don’t run volumetric ANIMAL segmentation -lobe_atlas <model> Use lobe atlas for ANIMAL segmentation (mandatory with -animal): “icbm152nl-VI” (ICBM152 generation VI symmetric model), “icbm152nl-2009a” (ICBM152 2009a symmetric model)
— Pipeline control ------------------------------------------------------------
-run Run the pipeline. -status-from-files Compute pipeline status from files -print-stages Print the pipeline stages. -print-status Print the status of each pipeline. -make-graph Create dot graph file. -make-filename-graph Create dot graph of filenames. -print-status-report Writes a CSV status report to file in cwd.
— Stage Control ---------------------------------------------------------------
-reset-all Start the pipeline from the beginning. -reset-from <stage_name> Restart from the specified stage. -reset-after <stage_name> Restart after the specified stage. -reset-to <stage_name> Run up to and including the specified stage. -reset-running Restart currently running jobs. [default] -no-reset-running opposite of -reset-running
Command-line Interface to CIVET
A command-line interface is available on some development systems:
USAGE (enumerating each subject ID in command line: <id1>, <id2> … <idn>):
CIVET_Processing_Pipeline -sourcedir <dir> -targetdir <dir> -prefix <prefix> [options] <id1> <id2> … <idn> -run > <logfile>
ALTERNATIVE USAGE (reading in subject list via text file: -id-file <idfile>):
CIVET_Processing_Pipeline -sourcedir <dir> -targetdir <dir> -prefix <prefix> [options] -id-file <idfile> -run > <logfile>
In any CIVET run, you must specify:
-
-sourcedir <dir>: directory of source images -
-targetdir <dir>: output directory -
-prefix <name>: the source images prefix (name of study) - a list of subjects to process (an enumeration or a list in a simple text file)
- an action command (like
-run). See the other action commands under Pipeline control options listed above.
Common additional options are:
-
-N3-distance <value>: spline distance for correction of non-uniformities using N3 (200 for 1.5T; 50 (old scans) or 100–125 (new scans) for 3.0T) -
-headheight <value>: height of head for neck-cropping to aid linear registration (about 170mm for young children, 175mm for adults) -
-lsq12: use affine 12-parameter transformation to stereotaxic space -
-resample-surfaces: resample the cortical surfaces, thus providing vertex-based areas and volumes as well as lobar segmentation
To find out about the CIVET stages in order to do a reset , first add the -print-stages option to your usual CIVET command to simply list the stages (it will print stages instead of running):
CIVET_Processing_Pipeline -sourcedir <dir> -targetdir <dir> -prefix <prefix> [options] -id-file <idfile> -run -print-stages > <logfile>
Then use one of the Stage Control options to restart CIVET relative to a given stage. You should rarely have to do this. One example:
CIVET_Processing_Pipeline -sourcedir <dir> -targetdir <dir> -prefix <prefix> [options] -id-file <idfile> -run -reset-from mincbet_mask_stx > <logfile>
The stage names are:
- clean_native_scan
- nuc_t1_native
- stx_register
- stx_tal_to_7
- stx_tal_to_6
- tal_t1
- nuc_inorm_t1
- mincbet_mask_stx
- nlfit
- mask_classify
- pve
- cortical_masking
- surface_classify
- create_wm_hemispheres
- extract_white_surface_left
- extract_white_surface_right
- laplace_field
- gray_surface_left
- mid_surface_left
- gray_surface_right
- mid_surface_right
- cls_volumes
- gyrification_index_left_gray
- verify_laplace
- gyrification_index_left_white
- verify_surfsurf
- surface_registration_left
- gyrification_index_right_gray
- cerebral_volume
- surface_fit_error
- gyrification_index_left_mid
- surface_registration_right
- surface_resample_left_white
- surface_resample_right_white
- verify_image
- gyrification_index_right_white
- surface_resample_left_mid
- surface_resample_left_gray
- thickness_tlink_30mm_right
- thickness_tlink_30mm_left
- surface_resample_right_mid
- surface_resample_right_gray
- gyrification_index_right_mid
- surface_area_rsl_40mm_left_mid
- verify_atlas
- resample_right_thickness_tlink_30mm
- resample_left_thickness_tlink_30mm
- asymmetry_map_position
- surface_area_rsl_40mm_right_mid
- surface_volume_rsl_40mm_right
- surface_volume_rsl_40mm_left
- asymmetry_rms_tlink_30mm
- lobe_thickness_left_30mm
- lobe_volume_left_40mm
- lobe_volume_right_40mm
- lobe_native_area_right
- lobe_thickness_right_30mm
- lobe_native_area_left
- verify_clasp
- lobe_area_left_40mm
- lobe_area_right_40mm
- verify_civet